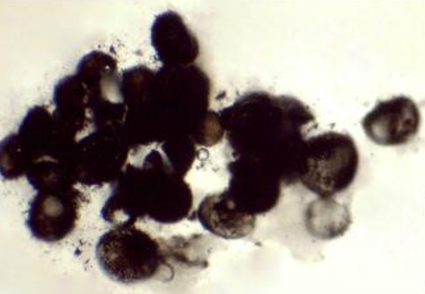
Abstract
A new species of Acrocalymma, A. guizhouense, isolated from the rhizosphere soil of Perilla frutescens, is introduced. Morphological characteristics, culture characteristics on PDA, OA, MEA, and phylogenetic analyses based on multi-locus datasets (SSU + LSU + ITS) support the establishment of the new species. Morphologically, A. guizhouense is distinguished from other species of Acrocalymma in having narrow conidia with a width of 1.5-2.5 µm. Phylogenetically, A. guizhouense formed a separated subclade with well-supported (100 MLBS/1.00 BYPP) value, which confirmed the taxonomic placement in the genus Acrocalymma.
References
Crous, P.W., Shivas, R.G., Quaedvlieg, W., Van der Bank, M., Zhang, Y., Summerell, B.A., Guarro, J., Wingfield, M.J., Wood, A.R., Alfenas, A.C., Braun, U., Cano-Lira, J.F., García, D., Marin-Felix, Y., Alvarado, P., Andrade, J.P., Armengol, J., Assefa, A., den Breeÿen, A., Camele, I., Cheewangkoon, R., De Souza, J.T., Duong, T.A., Esteve-Raventós, F., Fournier, J., Frisullo, S., García-Jiménez, J., Gardiennet, A., Gené, J., Hernández-Restrepo, M., Hirooka, Y., Hospenthal, D.R., King, A., Lechat, C., Lombard, L., Mang, S.M., Marbach, P.A.S., Marincowitz, S., Marin-Felix, Y., Montaño-Mata, N.J., Moreno, G., Perez, C.A., Pérez Sierra, A.M., Robertson, J.L., Roux, J., Rubio, E., Schumacher, R.K., Stchigel, A.M., Sutton, D.A., Tan, Y.P., Thompson, E.H., Vanderlinde, E., Walker, A.K., Walker, D.M., Wickes, B.L., Wong, P.T.W. & Groenewald, J.Z. (2014) Fungal Planet description sheets: 214–280. Persoonia-Molecular Phylogeny and Evolution of Fungi 32 (1): 184–306. https://doi.org/10.3767/003158514X682395
Dong, W., Wang, B., Hyde, K.D., McKenzie, E.H., Raja, H.A., Tanaka, K., Abdel-Wahab, M.A., Abdel-Aziz, F.A., Doilom, M., Phookamsak, R., Hongsanan, S., Wanasinghe, D.N., Yu, X., Wang, G., Yang, H., Yang, J., Thambugala, K.M., Tian, Q., Luo, Z., Yang, J., Miller, A.N., Fournier, J., Boonmee, S., Hu, D., Nalumpang, S. & Zhang, H. (2020) Freshwater Dothideomycetes. Fungal Diversity 105: 319–575. https://doi.org/10.1007/s13225-020-00463-5
Irwin, J.A.G. (1972) Stagonospora root and crown rot of lucerne. Australasian Plant Pathology 1 (4): 29–30. https://doi.org/10.1071/app9720029
Jayasiri, S.C., Hyde, K.D., Jones, E.B.G., McKenzie, E., Jeewon, R., Phillips, A.J.L., Bhat, D.J., Wanasinghe, D.N., Liu, J.J., Lu, Y., Kang, J., Xu, J. & Karunarathna, S.C. (2019) Diversity, morphology and molecular phylogeny of Dothideomycetes on decaying wild seed pods and fruits. Mycosphere 10 (1): 1–186. https:// doi.org/10.5943/mycosphere/10/1/1
Kalyaanamoorthy, S., Minh, B.Q., Wong, T.K.F., von Haeseler, A. & Jermiin, L.S. (2017) ModelFinder: fast model selection for accurate phylogenetic estimates. Nature Methods 14: 587–589. https://doi.org/10.1038/nmeth.4285
Katoh, K. & Standley, D.M. (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Molecular Biology and Evolution 30 (4): 772–780. https://doi.org/10.1093/molbev/mst010
Mortimer, P.E., Jeewon, R., Xu, J.C., Lumyong, S. & Wanasinghe, D.N. (2021) Morpho-phylo taxonomy of novel Dothideomycetous fungi associated with dead woody twigs in Yunnan Province, China. Frontiers in Microbiology 12: 654683. https://doi.10.3389/fmicb.2021.654683
Nguyen, L.T., Schmidt, H.A., von Haeseler, A. & Minh, B.Q. (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Molecular biology and evolution 32 (1): 268–274. https://doi.org/10.1093/molbev/ msu300
Ronquist, F., Teslenko, M., van der Mark, P., Ayres, D.L., Darling, A., Höhna, S., Larget, B., Liu, L., Suchard, M.A. & Huelsenbeck, J.P. (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Systematic Biology 61 (3): 539–542. https://doi.org/10.1093/sysbio/sys029
Shoemaker, R.A., Babcock, C.E. & Irwin, J.A.G. (1991) Massarina walkeri n.sp., the teleomorph of Acrocalymma medicaginis from Medicago sativa contrasted with Leptosphaeria pratensis, L. weimeri n. sp., and L. viridella. Canadian Journal of Botany 69 (3): 569–573. https://doi.org/10.1139/b91-077
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Molecular Biology and Evolution 30 (12): 2725–2729. https://doi.org/10.1093/molbev/mst197
Tennakoon, D.S., Kuo, C.H., Maharachchikumbura, S.S.N., Thambugala, K.M., Gentekaki, E., Phillips, A.L.J., Bhat, D.J., Wanasinghe, D.N., de Silva, N.I., Promputtha, I. & Hyde, K.D. (2021) Taxonomic and phylogenetic contributions to Celtis formosana, Ficus ampelas, F. septica, Macaranga tanarius and Morus australis leaf litter inhabiting microfungi. Fungal Diversity 108: 1–215. https://doi.org/10.1007/s13225-021-00474-w
Trakunyingcharoen, T., Lombard, L., Groenewald, J.Z., Cheewangkoon, R., Toanun, C., Alfenas, A.C. & Crous, P.W. (2014) Mycoparasitic species of Sphaerellopsis, and allied lichenicolous and other genera. IMA Fungus 5: 391–414. https://doi.org/10.5598/imafungus.2014.05.02.05
Vilgalys, R. & Hester, M. (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. Journal of Bacteriology 172: 4238–4246. https://doi.org/10.1128/jb.172.8.4238-4246.1990
White, T.J., Bruns, T., Lee, S. & Taylor, J. (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis, M.A., Gelfand, D.H., Sninsky, J.J. & White, T.J. (Eds.) PCR protocols: a guide to methods and applications, Academic Press, San Diego, California, pp 315–322.
Zhang, H., Hyde, K.D., Mckenzie, E.H., Bahkali, A.H. & Zhou, D. (2012) Sequence data reveals phylogenetic affinities of Acrocalymma aquatica sp. nov., Aquasubmersa mircensis gen. et sp. nov. and Clohesyomyces aquaticus (freshwater coelomycetes). Mycologie 33 (3): 333–346. https://doi.org/10.7872/crym.v33.iss3.2012.333
Zhang, Z.Y., Shao, Q.Y., Li, X., Chen, W.H., Liang, J.D., Han, Y.F., Huang, J.Z. & Liang, Z.Q. (2021) Culturable fungi from urban soils in China I: description of 10 new taxa. Microbiology Spectrum 9: e00867–00821. https://doi.org/10.1128/Spectrum.00867-21